Pass

The acronym PASS stands for Prediction of Activity Spectra for Substances. Using structural formula of a drug-like substance as an input, one obtains its estimated biological activity profile as an output. The predicted biological activity list includes the names of the probable activities with two probabilities: Pa – likelihood of belonging to the class of “Actives” and Pi – likelihood of belonging to the class of “Inactives”.
By default, all activities with Pa>Pi are considered as probable; however, depending on the particular tasks, the user may choose any other cutoff for selecting the probable “Actives”.
PASS has been well accepted by the research community, and is now actively used in the field of medicinal chemistry, by both academic organizations and pharma companies. There are over 1,200 publications described PASS approach and its applications. Overview on some papers is provided here.


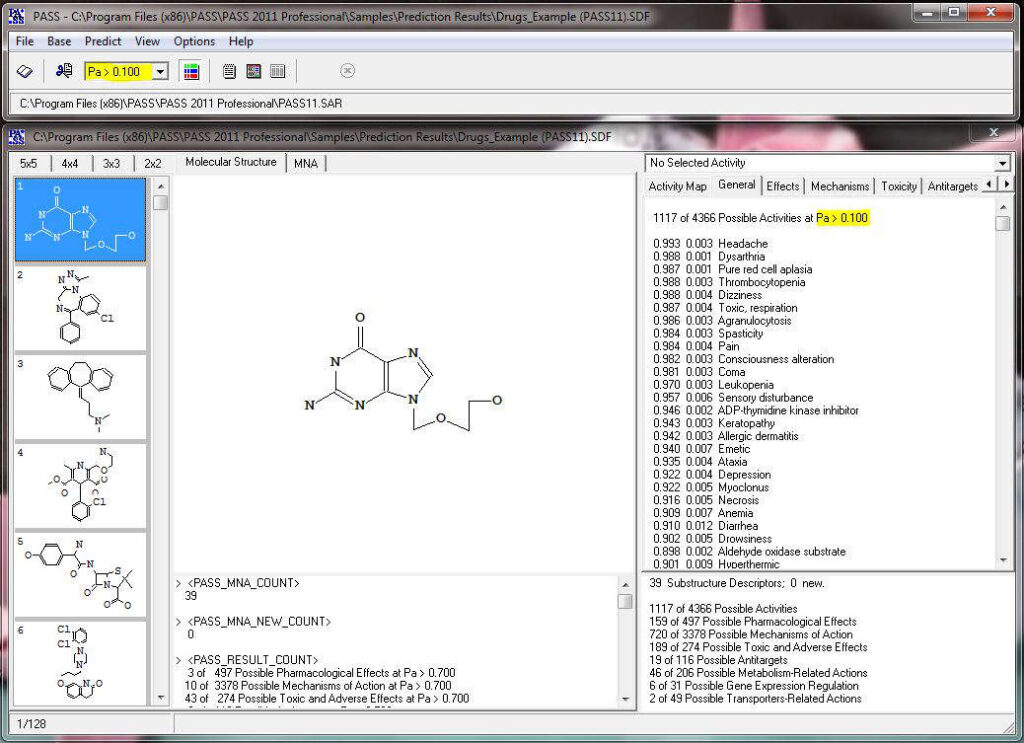
Activity prediction for a chemical substance by PASS
The slide show demonstrating the look-and-feel of PASS can be found here as well as on our Facebook site. A particularly useful tool to analyze and utilize PASS results further is PharmaExpert.
Key features of PASS 2022

In PASS 2022, the MNA descriptors (for prediction of activity spectra or for adding substances to SAR Base) are generated if structure corresponds to the following criteria:
- each of the atoms in a molecule must be presented by atom symbol from the periodic table. Symbols of unspecified atom A, Q, *, or R group labels are not allowed;
- each of the bonds in a molecule must be covalent bond presented by single, double or triple bond types only.
All other limitations on the structural formulae implemented in the previous PASS versions (only one uncharged component, minimum three carbon atoms in the structure, MW<1,250) are not applied anymore.
If the structure does not correspond to these criteria or the input data contains any other errors, a message about the first critical error will be received.
For a multicomponent structure, only the largest component (with the largest number of heavy atoms) is taken into account.
Based on the prediction results, you can evaluate the contribution of each of the atoms of the structure to the estimated biological activity. Select the desired biological activity in the predicted activity spectra by clicking on it; then, each of the atoms of the structure will be colored according to the following scheme:
Light Green Pa = 1, Pi = 0 (atom promotes activity)
Light Red Pa = 0, Pi = 1 (atom promotes inactivity)
Light Blue Pa = 0, Pi = 0 (atom does not generate any signal)
Grey Pa = 0.33, Pi = 0.33 (atom equivocal for weak signal)

Acyclovir, selected activity – “Antineoplastic enhancer”.
More about product
Price request
PASS packages
PASS Standard
Standard software package includes the standard SAR Base (Structure-Activity Relationship knowledgebase). Currently, standard version of PASS can predict 8,565 biological activities (1,957 activities are predicted by default).Depending on the particular purposes, the user may include into the predictable activity list any of the 8,565activity types using the “Selection” procedure.
PASS Refined
PASS Refined package provides all functions of PASS Standard and can predict 1,957 biological activities that are the most topical.
PASS Professional
PASS Professional package provides all functions of PASS Standard and the additional options to create, train and validate your proprietary SAR Base. With this package, you could make your own and unique SAR Base, and use it for further predictions in house on the exclusive basis. You may also add some additional data on structure-activity relationships to the standard SAR Base, and re-train the program to obtain the updated SAR Base. Thus, locally you would have a unique variant of PASS.
PASS Light
With PASS Light, you can create, train and validate your proprietary SAR base, and use it for further predictions. The standard SAR Base is not included.
PASS customized
According to your potential focus on particular types of biological activities, a customized version of PASS can be prepared, which can predict a restricted number of activities as per your selection. The command line utilities PASS 2022 to CSV, PASS 2022 to SDF, and PASS 2022 to TXT for use in pipelines are also available.
PASS + PharmaExpert package
Any of the product PASS Standard, PASS Refined, PASS Professional, PASS Light or PASS customized, can be ordered in a package with PharmaExpert.
Current statistics
PASS 2022 SAR Base is based on the information about structure-activity relationships of 1,614,066 substances; the total number of experimentally determined pairwise structure-activity records is 5,174,855. 8,565 biological activities can be predicted with average accuracy about 0.93; by default, 1,957 activities are predicted with average accuracy 0.97.

The basis > How it works
BIOLOGICAL ACTIVITY
Biological activity is the result of a chemical compound’s interaction with biological objects. It depends on the characteristics of the compound (structure of molecule), the biological object (kind, sex, age, etc.), details of the exposure (route of administration, dosage), and peculiarities of the experimental conditions. In PASS, biological activities are described qualitatively («active» or «inactive»).
BIOLOGICAL ACTIVITY SPECTRUM
The Biological Activity Spectrum of a chemical compound is the set of different types of biological activity that reflect the results of the compound’s interaction with various biological entities. It represents the “intrinsic” property of a compound depending only on its chemical structure. Though this may be a generalization, it provides the possibility for combining information from many different sources in the same training set, which is necessary because no one particular publication comprehensively covers all the various facets of the biological action of a compound.
CHEMICAL STRUCTURE REPRESENTATION
The 2D structural formulae of organic compound were chosen as the basis for the description of chemical structures, because this is the only information available at the early stage of research (compounds may only be designed but not synthesized yet). The structure descriptors we use, which we call «Multilevel Neighborhoods of Atoms» (MNA), were specifically designed for chemical structure representation for SAR analysis realized in PASS and similar approaches (Filimonov D. et al., 1999). LINK
Extended Connectivity Fingerprints (ECFP), which were developed later (Glen R.C. et al., 2006), are based on the same idea as MNA descriptors:

MNA descriptors, unlike ECFP, preserve the connectivity between atoms of different layers in the form of a nested bracket structure. They are based on the molecular structure representation, which includes all hydrogen atoms according to the valences and charges of atoms and does not specify bond types. Therefore, they inherently include the information about the type of hybridization of atomic orbitals.
EQUIVALENT STRUCTURES
The chemical compounds are considered equivalent in PASS if their molecular structures have the same MNA descriptors set. Since MNA descriptors do not represent the stereochemical peculiarities of a molecule, structures that only differ by stereochemistry are formally considered equivalent.
SAR BASE
The SAR Base contains vocabularies of MNA descriptors and activity names, the database of the substances’ structures represented by MNA descriptors, their biological activity spectrum, and data on the structure-activity relationships (SAR). Unfortunately, it is currently impossible to collect sufficiently large numbers of active compounds for all activities from available sources. This is why some activity types are represented in the general PASS training set by more than 300,000 drug-like compounds, while some others are only represented by a few ones. The supplied with PASS SAR Bases consists of the substances with at list one known activity.
PASS 2022 SAR Base is based on the information about structure-activity relationships obtained with an in-house training set of more than 1,614,066 compounds with 10,112 known biological activities. This training set is continuously curated and expanded. SAR Base can also be replaced by an exclusive knowledgebase, which can be created using in-house data. SAR Base together with the user-defined constraints of biological activities of interest and relevant parameters provides PASS the starting point for the computational prediction.
More detailed description of PASS method is available in several publications, for instance in the paper: Filimonov et al., 2014. LINK(PDF file may be obtained on request).
Competitive advantages and peculiarities
- Several thousand biological activities are predicted simultaneously, including: mechanisms of action, pharmacological effects, toxic and side effects, interaction with antitargets, interaction with drug metabolizing enzymes (inhibition, induction, interaction as a substrate), interaction with transporter protein (inhibition, stimulation, interaction as a substrate), changing gene expression of individual genes (increase, decrease). Due to such extensive analysis, the general pharmacological potential of molecules under study is disclosed.
- The user may select a particular pharmacotherapeutic field or a certain set of analyzed activities relevant to the purpose of the study. Such selection facilitates the analysis of the output information and reduce the computation time, which is particularly important in case of large chemical libraries.
- Algorithm of prediction is statistically robust with respect to the incompleteness of the data in the training set. Since no one biologically active compound is tested against all known biological activities, it is necessary to obtain the best possible estimations even using the imperfect information.
- The prediction is based on the structural formula of the compound that allows one to apply it to the virtual structures.
- Evaluation of the pharmaceutical substances’ action on many pharmacological targets to identify the hidden pharmacological potential of the launched pharmaceuticals, provides the ideas regarding possible drug repurposing.
- Prediction of the potential averse/toxic effects of drug-like compounds leads to their filtering out at the early stages of research.
- Re-training of the program using the customer’s training set lead to unique proprietary SAR bases creation, which can be used in house for further predictions.
- High rate of computation allows efficient multitarget virtual screening of large chemical libraries containing many millions structural formulae of drug-like compounds.
PASS applications
- Medicinal chemistry
- Computational chemistry
- Drug discovery / drug development
- Drug repositioning
- Chemical toxicity
- Safety assessment
- Pharmacogenomics, chemogenomics
- SAR (qualitative structure-activity relationship)
- Natural compound effects
- Translational research / translational medicine